orthodb
1、数据库

orthodb数据:
odb10v0_levels.tab.gz: NCBI taxonomy nodes where Ortho DB orthologous groups (OGs) are calculated
odb10v0_species.tab.gz: Ortho DB individual organism (aka species) ids based on NCBI taxonomy ids (mostly species level)
odb10v0_level2species.tab.gz: correspondence between level ids and species ids
odb10v0_genes.tab.gz: Ortho DB genes with some info
odb10v0_OGs.tab.gz: Ortho DB orthologous groups
odb10v0_OG2genes.tab.gz: OGs to genes correspondence
odb10v0_OG_xrefs.tab.gz: OG associations with GO, COG and InterPro ids
v9_v10_OGs_map.tab.gz mappings between the previous and current release orthologous group ids
odb10v0_fasta_<root>.tgz tar-ball with one fasta file per taxon id in the given root (bacteria,metazoa,fungi,plants)
2、odb10v0_levels.tab:
1. level NCBI taxonomy id
2. scientific name
3. total non-redundant count of genes in all underneath clustered species(在聚集的物种下面的所有的基因的总非重复计数)
4. total count of OGs built on it
5. total non-redundant count of species underneath

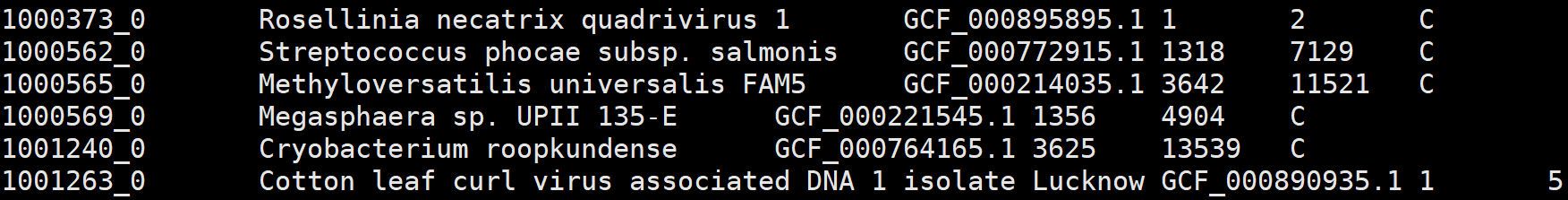
3、odb10v0_species.tab.gz
1. Ortho DB individual organism id, based on NCBI tax id
2. scientific name inherited from the most relevant NCBI tax id
3. genome asssembly id, when available
4. total count of clustered genes in this species
5. total count of the OGs it participates
6. mapping type, clustered(C) or mapped(M)
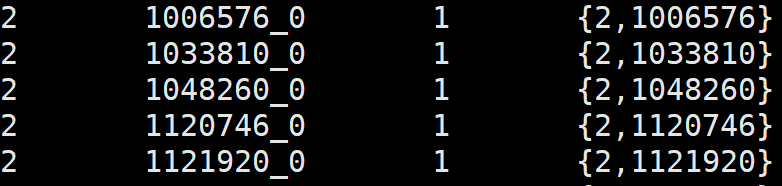
4、odb10v0_level2species.tab
1. top-most level NCBI tax id, one of [2,2157,2759,10239]
2. Ortho DB organism id
3. number of hops between the top-most level id and the NCBI tax id assiciated with the organism
4. ordered list of Ortho DB selected intermediate levels from the top-most level to the bottom one
5、odb10v0_genes.tab
1. Ortho DB unique gene id (not stable between releases)
2. organism tax id
3. protein original sequence id, as downloaded along with the sequence
4. Uniprot id, evaluated by mapping
5. ENSEMBL gene name, evaluated by mapping
6. NCBI gid, evaluated by mapping
7. description, evaluated by mapping
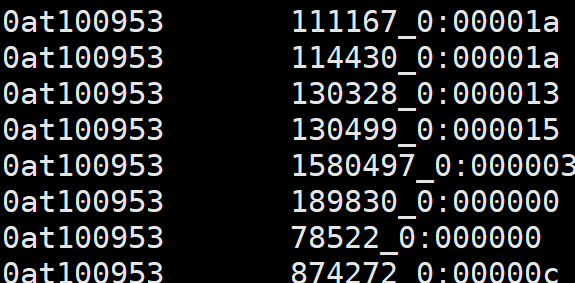
6、odb10v0_OG2genes.tab
1. OG unique id
2. Ortho DB gene id
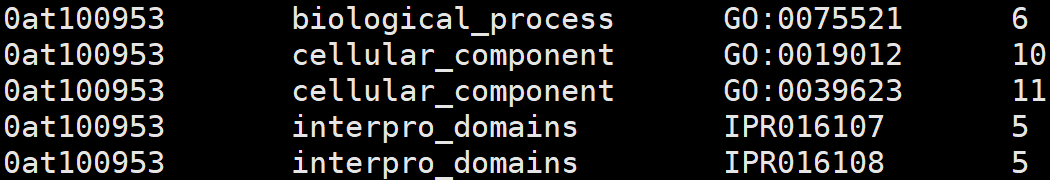
7、odb10v0_OG_xrefs.tab
1. OG unique id
2. external DB or DB section
3. external identifier
4. number of genes in the OG associated with the identifier
参考
https://www.orthodb.org/?page=filelist
orthodb的更多相关文章
- 【基因组预测】braker2基因结构注释要点记录
目录 流程使用 问题 记录下braker2的使用要点,以备忘记. 流程使用 braker2有很多流程,根据你的数据:组装的基因组.转录组.蛋白(同源,包括近缘或远缘)选择不同流程,官网有说明: htt ...
随机推荐
- 零基础学习python_字符串(14-15课)
今天回顾下我之前学习python的第一个对象——字符串,这个对象真蛋疼,因为方法是最多的,也是最常见的类型,没有之一... 内容有点多,我就搜了下网上的资料,转载下这个看起来还不错的网址吧:http: ...
- 网易云音乐mp3外链、真实地址下载方法
一个网易音乐外链地址长期有效,很简单的方法: 第一步打开网易云音乐,随便找到一首歌,播放,复制网址的ID, 例如:杨钰莹的心雨,网址是: http://music.163.com/#/song?id= ...
- 【Selenium-WebDriver自学】Log4J的设置(十五)
==================================================================================================== ...
- mysql之pymsql的使用
# -*- coding:utf-8 -*- import pymysql user = input('请输入用户名:') pwd = input('请输入密码:') # 1.连接 conn = py ...
- c++引用(修改引用的值)
当我们希望修改某个函数的返回值时,通常我们会返回这个值的引用(因为函数返回值其实是返回那个值得一份拷贝而已,所以想要修改必须使用引用): .h文件 #pragma once #include < ...
- C#打印0到100的素数
static void Main(string[] args) { //输出1-100的素数 bool res; ; ; i < ; i++) { res = true; ; j < i; ...
- 传输模型, tcp socket套接字
osi七层模型 tcp/ip四层模型 socket套接字 tcp 协议是可靠的 包括 三次握手 四次挥手 import socket # server server = socket.socket( ...
- fabric-ca1.03安装
前面的文档已经成功的安装了fabric1.0.1的e2e例子.之后代码换成1.0.3版本按步骤重新安装一下,就可以切换到1.0.3了.1.0.3的脚本和启动命令没有变化,还是用的1.0.1的. 1.准 ...
- scrapy工作原理探秘
def _next_request_from_scheduler(self, spider):#engine从调度器取得下一个request slot = self.slot request = sl ...
- ubuntu卸载福昕阅读器
在安装目录找到maintenancetool.sh运行之 ~/opt/foxitsoftware/foxitreader
