poj 1080 Human Gene Functions(lcs,较难)
| Time Limit: 1000MS | Memory Limit: 10000K | |
| Total Submissions: 19573 | Accepted: 10919 |
Description
A human gene can be identified through a series of time-consuming biological experiments, often with the help of computer programs. Once a sequence of a gene is obtained, the next job is to determine its function.
One of the methods for biologists to use in determining the function of a new gene sequence that they have just identified is to search a database with the new gene as a query. The database to be searched stores many gene sequences and their functions – many researchers have been submitting their genes and functions to the database and the database is freely accessible through the Internet.
A database search will return a list of gene sequences from the database that are similar to the query gene.
Biologists assume that sequence similarity often implies functional similarity. So, the function of the new gene might be one of the functions that the genes from the list have. To exactly determine which one is the right one another series of biological experiments will be needed.
Your job is to make a program that compares two genes and determines their similarity as explained below. Your program may be used as a part of the database search if you can provide an efficient one.
Given two genes AGTGATG and GTTAG, how similar are they? One of the methods to measure the similarity
of two genes is called alignment. In an alignment, spaces are inserted, if necessary, in appropriate positions of
the genes to make them equally long and score the resulting genes according to a scoring matrix.
For example, one space is inserted into AGTGATG to result in AGTGAT-G, and three spaces are inserted into GTTAG to result in –GT--TAG. A space is denoted by a minus sign (-). The two genes are now of equal
length. These two strings are aligned:
AGTGAT-G
-GT--TAG
In this alignment, there are four matches, namely, G in the second position, T in the third, T in the sixth, and G in the eighth. Each pair of aligned characters is assigned a score according to the following scoring matrix. 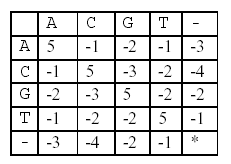
denotes that a space-space match is not allowed. The score of the alignment above is (-3)+5+5+(-2)+(-3)+5+(-3)+5=9.
Of course, many other alignments are possible. One is shown below (a different number of spaces are inserted into different positions):
AGTGATG
-GTTA-G
This alignment gives a score of (-3)+5+5+(-2)+5+(-1) +5=14. So, this one is better than the previous one. As a matter of fact, this one is optimal since no other alignment can have a higher score. So, it is said that the
similarity of the two genes is 14.
Input
Output
Sample Input
2
7 AGTGATG
5 GTTAG
7 AGCTATT
9 AGCTTTAAA
Sample Output
14
21
题目大意是就是求两基因的相似度,先要在每个基因对中加入若干空格,然后再依次加上匹配度,详见上表,则相似度就是最大的匹配度和
例如对于测试数据一,加上空格则变成
AGTGAT--G
-GT----TAG
则相似度就是(-3)+5+5+(-2)+5+(-1)+5=14,可以证明这是最大的,故为所求
此题为dp,详见代码
#include<iostream>
#include<cstdlib>
#include<cstdio>
#include<cstring>
#include<algorithm>
#include<cmath>
using namespace std;
int map[][]={ {,-,-,-,-},
{-,,-,-,-},
{-,-,,-,-},
{-,-,-,,-},
{-,-,-,-,} };
char str1[];
char str2[];
int f[];
bool vis[][];
int ans[][];
int DFS ( int x , int y )
{ int i ;
int xy=;
if ( x == - )
{
for ( i = ; i <= y ; i ++ )
xy += map[str2[i]][] ;
return xy ;
}
if ( y == - )
{
for ( i = ; i <= x ; i ++ )
xy += map[str1[i]][] ;
return xy ;
}
if ( vis[x][y] )
return ans[x][y] ;
vis[x][y]=true;
ans[x][y] = max ( DFS ( x - , y - ) + map[str1[x]][str2[y]] , max ( DFS(x,y-)+map[][str2[y]] , DFS(x-,y)+map[str1[x]][] )) ;
return ans[x][y] ;
}
int main()
{
f['A']=;
f['C']=;
f['-']=;
f['G']=;
f['T']=;
int t ;
cin >> t ;
while ( t -- )
{
int i , j ;
int len1 , len2 ;
cin >> len1 >> str1 ;
for ( i = ; i < len1 ; i ++ )
str1[i]=f[str1[i]]; cin >> len2 >> str2 ;
for ( i = ; i < len2 ; i ++ )
str2[i]=f[str2[i]];
memset ( vis , , sizeof ( vis ) ) ;
cout << DFS(len1-,len2-) << endl ;
}
return ;
}
#include<cstdio>
#include<cstdlib>
#include<iostream>
#include<algorithm> using namespace std;
/*dp,poj1080*/ int dp[][];//动态规划数据存放
int map[][];//用来存放原始数据 void map_init()
{
map['A']['A']=map['C']['C']=map['G']['G']=map['T']['T']=;
map['A']['C']=map['C']['A']=map['A']['T']=map['T']['A']=map['T'][' ']=map[' ']['T']=-;
map['A']['G']=map['G']['A']=map['C']['T']=map['T']['C']=map['G']['T']=map['T']['G']=map['G'][' ']=map[' ']['G']=-;
map['A'][' ']=map[' ']['A']=map['G']['C']=map['C']['G']=-;
map['C'][' ']=map[' ']['C']=-;
} int max_X3(int a,int b,int c)
{
if(a>b)
{
if(a>c)
return a;
else
return c;
}
else
{
if(b>c)
return b;
else
return c;
}
} int main()
{
int y;//全局次数
int i,j;//循环变量
int a,b;//用户输入
char str1[];
char str2[]; //初始化
map_init(); cin>>y;
while (y--)
{
scanf("%d %s",&a,str1);
scanf("%d %s",&b,str2); //初始化第一行第一列
dp[][]=;
for (i = ; i < a; i++)
dp[][i+] = dp[][i] + map[str1[i]][' ']; for (j = ; j < b; j++)
dp[j+][] = dp[j][] + map[str2[j]][' ']; for (i = ; i <= a; i++)
{
for (j = ; j <= b; j++)
{
dp[j][i] = max_X3(dp[j-][i-]+map[str2[j-]][str1[i-]],
dp[j-][i]+map[str2[j-]][' '],
dp[j][i-]+map[str1[i-]][' ']);
}
} cout<<dp[b][a]<<endl;
}
return ;
}
poj 1080 Human Gene Functions(lcs,较难)的更多相关文章
- poj 1080 ——Human Gene Functions——————【最长公共子序列变型题】
Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K Total Submissions: 17805 Accepted: ...
- POJ 1080 Human Gene Functions -- 动态规划(最长公共子序列)
题目地址:http://poj.org/problem?id=1080 Description It is well known that a human gene can be considered ...
- poj 1080 Human Gene Functions(dp)
题目:http://poj.org/problem?id=1080 题意:比较两个基因序列,测定它们的相似度,将两个基因排成直线,如果需要的话插入空格,使基因的长度相等,然后根据那个表格计算出相似度. ...
- dp poj 1080 Human Gene Functions
题目链接: http://poj.org/problem?id=1080 题目大意: 给两个由A.C.T.G四个字符组成的字符串,可以在两串中加入-,使得两串长度相等. 每两个字符匹配时都有个值,求怎 ...
- POJ 1080 Human Gene Functions
题意:给两个DNA序列,在这两个DNA序列中插入若干个'-',使两段序列长度相等,对应位置的两个符号的得分规则给出,求最高得分. 解法:dp.dp[i][j]表示第一个字符串s1的前i个字符和第二个字 ...
- POJ 1080 Human Gene Functions 【dp】
题目大意:每次给出两个碱基序列(包含ATGC的两个字符串),其中每一个碱基与另一串中碱基如果配对或者与空串对应会有一个分数(可能为负),找出一种方式使得两个序列配对的分数最大 思路:字符串动态规划的经 ...
- poj 1080 Human Gene Functions (最长公共子序列变形)
题意:有两个代表基因序列的字符串s1和s2,在两个基因序列中通过添加"-"来使得两个序列等长:其中每对基因匹配时会形成题中图片所示匹配值,求所能得到的总的最大匹配值. 题解:这题运 ...
- POJ 1080:Human Gene Functions LCS经典DP
Human Gene Functions Time Limit: 1000MS Memory Limit: 10000K Total Submissions: 18007 Accepted: ...
- POJ1080 Human Gene Functions(LCS)
题目链接. 分析: 和 LCS 差不多. #include <iostream> #include <cstdio> #include <cstdlib> #inc ...
随机推荐
- 关于工作与生活——HP大中华区总裁孙振耀撰文谈退休并畅谈人生
转自:http://blog.csdn.net/adaptiver/article/details/7494121 我有个有趣的观察,外企公司多的是25-35岁的白领, 40岁以上的员工很少,二三十岁 ...
- 【Navicat Premium】之连接Oracle数据库
1.首先,在连接之前,需要下载oracle官网提供的instantclient-basic-win32-11.2.0.1.0.zip包 官网:http://www.oracle.com/technet ...
- [转]python 书籍推荐
原地址: http://python.jobbole.com/85620/ https://github.com/jobbole/awesome-python-books http://blog.cs ...
- 多媒体开发之---开源库ffmeg的log之子解析
用了ffmeg快两年了,对其中的log甚是感兴趣,今天在做8148项目是,解读h264结构,看了<毕-新一代视频压缩编码标准h246> ,在第六章中的重排序里面看到了好熟悉的4x4矩阵zi ...
- java解析字符串拆分单独元素
有时候,需求要求传递多个字符串参数,但是方法参数已经固定为单个String,笔者在学习unity和android之间的消息传递时就遇到这个问题,所以就写了这么一个解析字符串拆分单独元素的方法. 示例: ...
- onCreate中获得控件的大小
@Override public void onCreate(Bundle savedInstanceState) { super.onCreate(savedInstanceState); setC ...
- LCD驱动程序(二)
上节我们主要是对fb_info结构体的配置,对fb_info结构体的配置主要分为一下步骤: static int lcd_init(void){ /* 1. 分配一个fb_info */ s3c_lc ...
- 【CodeM初赛A轮】D 分解质因数+暴力
题目描述树链是指树里的一条路径.美团外卖的形象代言人袋鼠先生最近在研究一个特殊的最长树链问题.现在树中的每个点都有一个正整数值,他想在树中找出最长的树链,使得这条树链上所有对应点的值的最大公约数大于1 ...
- Mybatis的动态SQL实现
一.动态SQL简介 MyBatis的强大特性之一便是它的动态 SQL.如果你有使用 JDBC 或其他类似框架的经验,你就能体会到根据不同条件拼接 SQL 语句有多么痛苦.拼接的时候要确保不能忘了必要的 ...
- JAVA面试题——JAVA编程题1(2015.07.22)
实现代码很简单: package com.xiaozan.shopping; import java.util.Arrays; public class ShoppingCart { ...
