[占位-未完成]scikit-learn一般实例之十二:用于RBF核的显式特征映射逼近
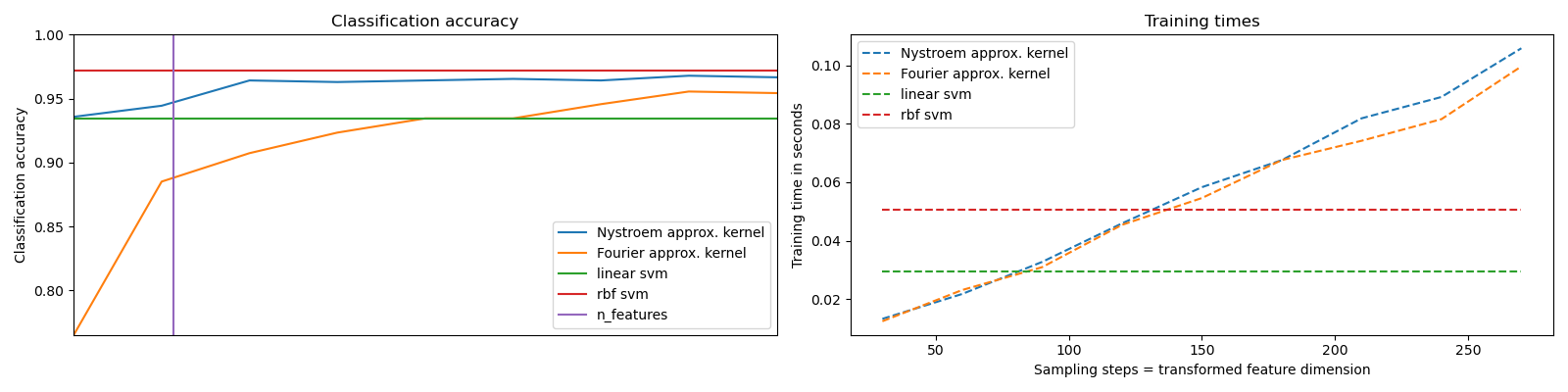
It shows how to use RBFSampler and Nystroem to approximate the feature map of an RBF kernel for classification with an SVM on the digits dataset. Results using a linear SVM in the original space, a linear SVM using the approximate mappings and using a kernelized SVM are compared. Timings and accuracy for varying amounts of Monte Carlo samplings (in the case of RBFSampler, which uses random Fourier features) and different sized subsets of the training set (for Nystroem) for the approximate mapping are shown.
Please note that the dataset here is not large enough to show the benefits of kernel approximation, as the exact SVM is still reasonably fast.
Sampling more dimensions clearly leads to better classification results, but comes at a greater cost. This means there is a tradeoff between runtime and accuracy, given by the parameter n_components. Note that solving the Linear SVM and also the approximate kernel SVM could be greatly accelerated by using stochastic gradient descent via sklearn.linear_model.SGDClassifier. This is not easily possible for the case of the kernelized SVM.
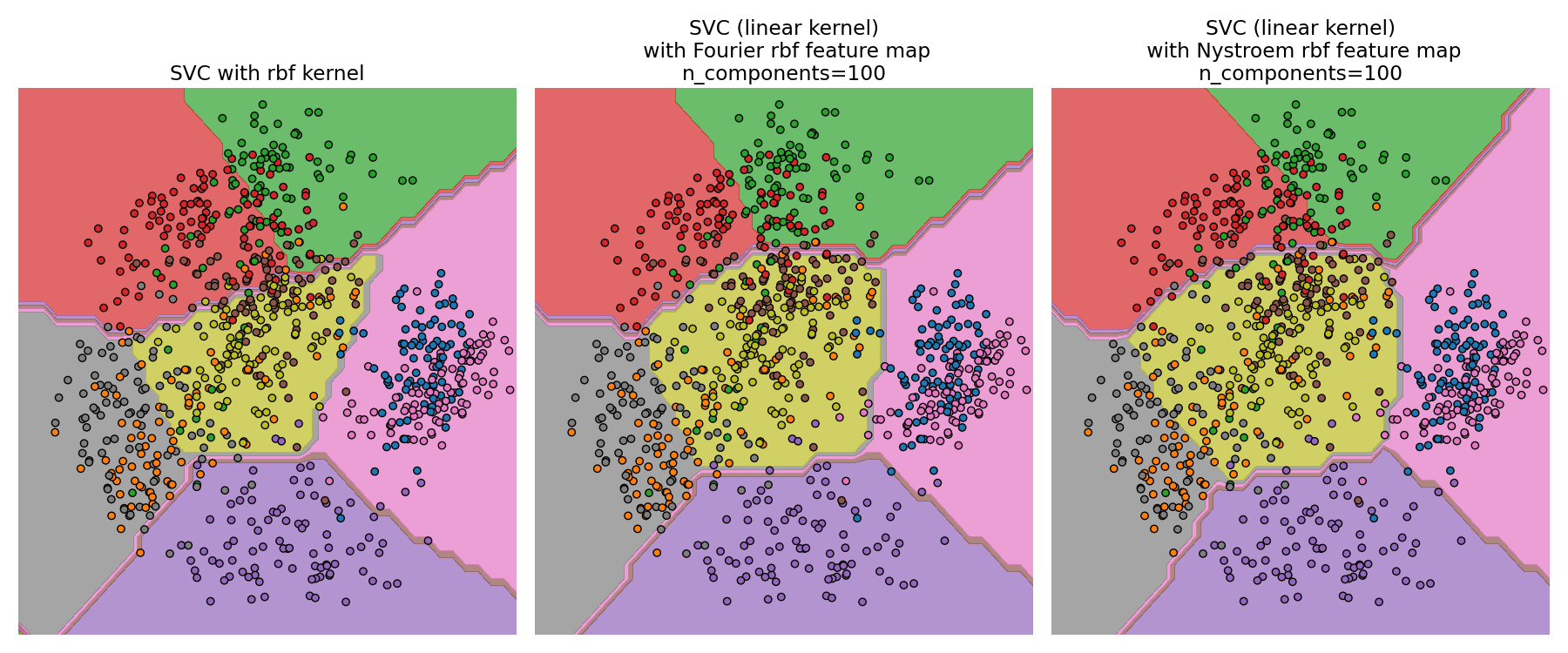
The second plot visualized the decision surfaces of the RBF kernel SVM and the linear SVM with approximate kernel maps. The plot shows decision surfaces of the classifiers projected onto the first two principal components of the data. This visualization should be taken with a grain of salt since it is just an interesting slice through the decision surface in 64 dimensions. In particular note that a datapoint (represented as a dot) does not necessarily be classified into the region it is lying in, since it will not lie on the plane that the first two principal components span.
The usage of RBFSampler and Nystroem is described in detail in Kernel Approximation.
print(__doc__)
# Author: Gael Varoquaux <gael dot varoquaux at normalesup dot org>
# Andreas Mueller <amueller@ais.uni-bonn.de>
# License: BSD 3 clause
# Standard scientific Python imports
import matplotlib.pyplot as plt
import numpy as np
from time import time
# Import datasets, classifiers and performance metrics
from sklearn import datasets, svm, pipeline
from sklearn.kernel_approximation import (RBFSampler,
Nystroem)
from sklearn.decomposition import PCA
# The digits dataset
digits = datasets.load_digits(n_class=9)
# To apply an classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.data)
data = digits.data / 16.
data -= data.mean(axis=0)
# We learn the digits on the first half of the digits
data_train, targets_train = data[:n_samples / 2], digits.target[:n_samples / 2]
# Now predict the value of the digit on the second half:
data_test, targets_test = data[n_samples / 2:], digits.target[n_samples / 2:]
#data_test = scaler.transform(data_test)
# Create a classifier: a support vector classifier
kernel_svm = svm.SVC(gamma=.2)
linear_svm = svm.LinearSVC()
# create pipeline from kernel approximation
# and linear svm
feature_map_fourier = RBFSampler(gamma=.2, random_state=1)
feature_map_nystroem = Nystroem(gamma=.2, random_state=1)
fourier_approx_svm = pipeline.Pipeline([("feature_map", feature_map_fourier),
("svm", svm.LinearSVC())])
nystroem_approx_svm = pipeline.Pipeline([("feature_map", feature_map_nystroem),
("svm", svm.LinearSVC())])
# fit and predict using linear and kernel svm:
kernel_svm_time = time()
kernel_svm.fit(data_train, targets_train)
kernel_svm_score = kernel_svm.score(data_test, targets_test)
kernel_svm_time = time() - kernel_svm_time
linear_svm_time = time()
linear_svm.fit(data_train, targets_train)
linear_svm_score = linear_svm.score(data_test, targets_test)
linear_svm_time = time() - linear_svm_time
sample_sizes = 30 * np.arange(1, 10)
fourier_scores = []
nystroem_scores = []
fourier_times = []
nystroem_times = []
for D in sample_sizes:
fourier_approx_svm.set_params(feature_map__n_components=D)
nystroem_approx_svm.set_params(feature_map__n_components=D)
start = time()
nystroem_approx_svm.fit(data_train, targets_train)
nystroem_times.append(time() - start)
start = time()
fourier_approx_svm.fit(data_train, targets_train)
fourier_times.append(time() - start)
fourier_score = fourier_approx_svm.score(data_test, targets_test)
nystroem_score = nystroem_approx_svm.score(data_test, targets_test)
nystroem_scores.append(nystroem_score)
fourier_scores.append(fourier_score)
# plot the results:
plt.figure(figsize=(8, 8))
accuracy = plt.subplot(211)
# second y axis for timeings
timescale = plt.subplot(212)
accuracy.plot(sample_sizes, nystroem_scores, label="Nystroem approx. kernel")
timescale.plot(sample_sizes, nystroem_times, '--',
label='Nystroem approx. kernel')
accuracy.plot(sample_sizes, fourier_scores, label="Fourier approx. kernel")
timescale.plot(sample_sizes, fourier_times, '--',
label='Fourier approx. kernel')
# horizontal lines for exact rbf and linear kernels:
accuracy.plot([sample_sizes[0], sample_sizes[-1]],
[linear_svm_score, linear_svm_score], label="linear svm")
timescale.plot([sample_sizes[0], sample_sizes[-1]],
[linear_svm_time, linear_svm_time], '--', label='linear svm')
accuracy.plot([sample_sizes[0], sample_sizes[-1]],
[kernel_svm_score, kernel_svm_score], label="rbf svm")
timescale.plot([sample_sizes[0], sample_sizes[-1]],
[kernel_svm_time, kernel_svm_time], '--', label='rbf svm')
# vertical line for dataset dimensionality = 64
accuracy.plot([64, 64], [0.7, 1], label="n_features")
# legends and labels
accuracy.set_title("Classification accuracy")
timescale.set_title("Training times")
accuracy.set_xlim(sample_sizes[0], sample_sizes[-1])
accuracy.set_xticks(())
accuracy.set_ylim(np.min(fourier_scores), 1)
timescale.set_xlabel("Sampling steps = transformed feature dimension")
accuracy.set_ylabel("Classification accuracy")
timescale.set_ylabel("Training time in seconds")
accuracy.legend(loc='best')
timescale.legend(loc='best')
# visualize the decision surface, projected down to the first
# two principal components of the dataset
pca = PCA(n_components=8).fit(data_train)
X = pca.transform(data_train)
# Generate grid along first two principal components
multiples = np.arange(-2, 2, 0.1)
# steps along first component
first = multiples[:, np.newaxis] * pca.components_[0, :]
# steps along second component
second = multiples[:, np.newaxis] * pca.components_[1, :]
# combine
grid = first[np.newaxis, :, :] + second[:, np.newaxis, :]
flat_grid = grid.reshape(-1, data.shape[1])
# title for the plots
titles = ['SVC with rbf kernel',
'SVC (linear kernel)\n with Fourier rbf feature map\n'
'n_components=100',
'SVC (linear kernel)\n with Nystroem rbf feature map\n'
'n_components=100']
plt.tight_layout()
plt.figure(figsize=(12, 5))
# predict and plot
for i, clf in enumerate((kernel_svm, nystroem_approx_svm,
fourier_approx_svm)):
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
plt.subplot(1, 3, i + 1)
Z = clf.predict(flat_grid)
# Put the result into a color plot
Z = Z.reshape(grid.shape[:-1])
plt.contourf(multiples, multiples, Z, cmap=plt.cm.Paired)
plt.axis('off')
# Plot also the training points
plt.scatter(X[:, 0], X[:, 1], c=targets_train, cmap=plt.cm.Paired)
plt.title(titles[i])
plt.tight_layout()
plt.show()
[占位-未完成]scikit-learn一般实例之十二:用于RBF核的显式特征映射逼近的更多相关文章
- C++面向对象类的实例题目十二
题目描述: 写一个程序计算正方体.球体和圆柱体的表面积和体积 程序代码: #include<iostream> #define PAI 3.1415 using namespace std ...
- 数据可视化实例(十二): 发散型条形图 (matplotlib,pandas)
https://datawhalechina.github.io/pms50/#/chapter10/chapter10 如果您想根据单个指标查看项目的变化情况,并可视化此差异的顺序和数量,那么散型条 ...
- Java开发笔记(七十二)Java8新增的流式处理
通过前面几篇文章的学习,大家应能掌握几种容器类型的常见用法,对于简单的增删改和遍历操作,各容器实例都提供了相应的处理方法,对于实际开发中频繁使用的清单List,还能利用Arrays工具的asList方 ...
- 框架源码系列十二:Mybatis源码之手写Mybatis
一.需求分析 1.Mybatis是什么? 一个半自动化的orm框架(Object Relation Mapping). 2.Mybatis完成什么工作? 在面向对象编程中,我们操作的都是对象,Myba ...
- [占位-未完成]scikit-learn一般实例之十:核岭回归和SVR的比较
[占位-未完成]scikit-learn一般实例之十:核岭回归和SVR的比较
- [占位-未完成]scikit-learn一般实例之十一:异构数据源的特征联合
[占位-未完成]scikit-learn一般实例之十一:异构数据源的特征联合 Datasets can often contain components of that require differe ...
- scikit learn 模块 调参 pipeline+girdsearch 数据举例:文档分类 (python代码)
scikit learn 模块 调参 pipeline+girdsearch 数据举例:文档分类数据集 fetch_20newsgroups #-*- coding: UTF-8 -*- import ...
- Scikit Learn: 在python中机器学习
转自:http://my.oschina.net/u/175377/blog/84420#OSC_h2_23 Scikit Learn: 在python中机器学习 Warning 警告:有些没能理解的 ...
- 《Android群英传》读书笔记 (5) 第十一章 搭建云端服务器 + 第十二章 Android 5.X新特性详解 + 第十三章 Android实例提高
第十一章 搭建云端服务器 该章主要介绍了移动后端服务的概念以及Bmob的使用,比较简单,所以略过不总结. 第十三章 Android实例提高 该章主要介绍了拼图游戏和2048的小项目实例,主要是代码,所 ...
随机推荐
- 用scikit-learn学习BIRCH聚类
在BIRCH聚类算法原理中,我们对BIRCH聚类算法的原理做了总结,本文就对scikit-learn中BIRCH算法的使用做一个总结. 1. scikit-learn之BIRCH类 在scikit-l ...
- 【C#附源码】数据库文档生成工具支持(Excel+Html)
[2015] 很多时候,我们在生成数据库文档时,使用某些工具,可效果总不理想,不是内容不详细,就是表现效果一般般.很多还是word.html的.看着真是别扭.本人习惯用Excel,所以闲暇时,就简单的 ...
- Servlet监听器笔记总结
监听器Listener的概念 监听器的概念很好理解,顾名思义,就是监视目标动作或状态的变化,目标一旦状态发生变化或者有动作,则立马做出反应. Servlet中的也有实现监听器的机制,就是Listene ...
- golang语言构造函数
1.构造函数定义 构造函数 ,是一种特殊的方法.主要用来在创建对象时初始化对象, 即为对象成员变量赋初始值,总与new运算符一起使用在创建对象的语句中.特别的一个类可以有多个构造函数 ,可根据其参数个 ...
- 负载均衡——nginx理论
nginx是什么? nginx是一个强大的web服务器软件,用于处理高并发的http请求和作为反向代理服务器做负载均衡.具有高性能.轻量级.内存消耗少,强大的负载均衡能力等优势. nginx架构? ...
- 15个C++项目列表
实验楼上有很多C++的实战项目,从简单到进阶,学习每个项目都可以掌握相应的知识点. 如果你还是C++新手的话,那么这个C++的项目列表你可以拿去练手实战开发,毕竟学编程动手实践是少不了的! 如果你不知 ...
- 设计模式之工厂模式VS抽象工厂
一.工厂模式主要是为创建对象提供过渡接口,以便将创建对象的具体过程屏蔽隔离起来,达到提高灵活性的目的. 工厂模式在<Java与模式>中分为三类:1)简单工厂模式(Simple Factor ...
- 远程连接mysql 1130错误解决方法
- firebug不能加载JS文件 ,无法进行JS脚本调试
提示: 本页面不包含 Javascript 如果 <script> 标签有 "type" 属性,其值应为 "text/javascript" 或者& ...
- 把int*传值给char*,打印出错误的数字
首先进入debug模式查看i的地址也就是ptr的值 以16进制位小端模式存储(一个整型四个字节,8位16进制数)(根据系统位数情况) 紧接着因为ptr是char*型指针变量,读取数据时按照一个字节一个 ...
