R生存分析AFT
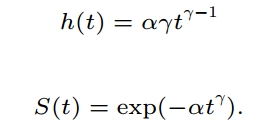
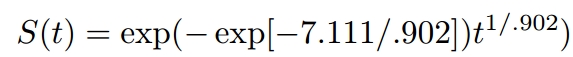
γ = 1/scale =1/0.902
α = exp(−(Intercept)γ)=exp(-(7.111)*γ)
> library(survival)
> myfit=survreg(Surv(futime, fustat)~1 , ovarian, dist="weibull",scale=0)
> summary(myfit) Call:
survreg(formula = Surv(futime, fustat) ~ 1, data = ovarian, dist = "weibull",
scale = 0)
Value Std. Error z p
(Intercept) 7.111 0.293 24.292 2.36e-130
Log(scale) -0.103 0.254 -0.405 6.86e-01 Scale= 0.902 Weibull distribution
Loglik(model)= -98 Loglik(intercept only)= -98
Number of Newton-Raphson Iterations: 5
n= 26
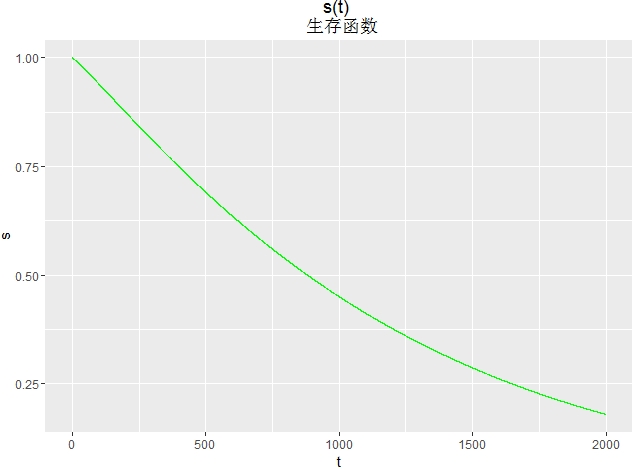
画生存函数图
d<- seq(0, 2000, length.out=10000) h<-1-pweibull(d,shape=1/0.902,scale=exp(7.111)) df<-data.frame(t=d,s=h) library(ggplot2) ggplot(df,aes(x=t,y=s))+
geom_line(colour="green")+
ggtitle("s(t) \n 生存函数")
1. Surv
Description
创建一个生存对象,通常用作模型公式中的响应变量。 参数匹配是此功能的特殊功能,请参阅下面的详细信息。
Surv(time, time2, event,
type=c('right', 'left', 'interval', 'counting', 'interval2', 'mstate'),
origin=0)
is.Surv(x)
Arguments
time
对于右删失数据,这是一个跟踪时间。对于区间数据,第一个参数是区间的开始时间。
event
状态指示,通常,0=活着,1=死亡。其他选择是TRUE/FALSE (TRUE = 死亡) or 1/2 (2=死亡)。对于区间删失数据,状态指示,0=右删失, 1=事件时间, 2=左删失, 3=区间删失.
右删失(Right Censoring):只知道实际寿命大于某数;
左删失(Left Censoring):只知道实际寿命小于某数;
区间删失(Interval Censoring):只知道实际寿命在一个时间区间内。
time2
区间删失区间的结束时间或仅对过程数据进行计数。
type
指定删失类型。 "right", "left", "counting", "interval", "interval2" or "mstate".
如果event变量是一个因子,假定type="mstate"。如果没有指定参数time2,type="right";如果指定参数time2,type="counting"
Surv使用示例
> str(lung)
'data.frame': 228 obs. of 10 variables:
$ inst : num 3 3 3 5 1 12 7 11 1 7 ...
$ time : num 306 455 1010 210 883 ...
$ status : num 2 2 1 2 2 1 2 2 2 2 ...
$ age : num 74 68 56 57 60 74 68 71 53 61 ...
$ sex : num 1 1 1 1 1 1 2 2 1 1 ...
$ ph.ecog : num 1 0 0 1 0 1 2 2 1 2 ...
$ ph.karno : num 90 90 90 90 100 50 70 60 70 70 ...
$ pat.karno: num 100 90 90 60 90 80 60 80 80 70 ...
$ meal.cal : num 1175 1225 NA 1150 NA ...
$ wt.loss : num NA 15 15 11 0 0 10 1 16 34 ...
> with(lung, Surv(time, status))
[1] 306 455 1010+ 210 883 1022+ 310 361 218
[10] 166 170 654 728 71 567 144 613 707
[19] 61 88 301 81 624 371 394 520 574
[28] 118 390 12 473 26 533 107 53 122
[37] 814 965+ 93 731 460 153 433 145 583
[46] 95 303 519 643 765 735 189 53 246
[55] 689 65 5 132 687 345 444 223 175
[64] 60 163 65 208 821+ 428 230 840+ 305
[73] 11 132 226 426 705 363 11 176 791
[82] 95 196+ 167 806+ 284 641 147 740+ 163
[91] 655 239 88 245 588+ 30 179 310 477
[100] 166 559+ 450 364 107 177 156 529+ 11
[109] 429 351 15 181 283 201 524 13 212
[118] 524 288 363 442 199 550 54 558 207
[127] 92 60 551+ 543+ 293 202 353 511+ 267
[136] 511+ 371 387 457 337 201 404+ 222 62
[145] 458+ 356+ 353 163 31 340 229 444+ 315+
[154] 182 156 329 364+ 291 179 376+ 384+ 268
[163] 292+ 142 413+ 266+ 194 320 181 285 301+
[172] 348 197 382+ 303+ 296+ 180 186 145 269+
[181] 300+ 284+ 350 272+ 292+ 332+ 285 259+ 110
[190] 286 270 81 131 225+ 269 225+ 243+ 279+
[199] 276+ 135 79 59 240+ 202+ 235+ 105 224+
[208] 239 237+ 173+ 252+ 221+ 185+ 92+ 13 222+
[217] 192+ 183 211+ 175+ 197+ 203+ 116 188+ 191+
[226] 105+ 174+ 177+ > str(heart)
'data.frame': 172 obs. of 8 variables:
$ start : num 0 0 0 1 0 36 0 0 0 51 ...
$ stop : num 50 6 1 16 36 39 18 3 51 675 ...
$ event : num 1 1 0 1 0 1 1 1 0 1 ...
$ age : num -17.16 3.84 6.3 6.3 -7.74 ...
$ year : num 0.123 0.255 0.266 0.266 0.49 ...
$ surgery : num 0 0 0 0 0 0 0 0 0 0 ...
$ transplant: Factor w/ 2 levels "0","1": 1 1 1 2 1 2 1 1 1 2 ...
$ id : num 1 2 3 3 4 4 5 6 7 7 ...
> Surv(heart$start, heart$stop, heart$event)
[1] ( 0.0, 50.0] ( 0.0, 6.0] ( 0.0, 1.0+]
[4] ( 1.0, 16.0] ( 0.0, 36.0+] ( 36.0, 39.0]
[7] ( 0.0, 18.0] ( 0.0, 3.0] ( 0.0, 51.0+]
[10] ( 51.0, 675.0] ( 0.0, 40.0] ( 0.0, 85.0]
[13] ( 0.0, 12.0+] ( 12.0, 58.0] ( 0.0, 26.0+]
[16] ( 26.0, 153.0] ( 0.0, 8.0] ( 0.0, 17.0+]
[19] ( 17.0, 81.0] ( 0.0, 37.0+] ( 37.0,1387.0]
[22] ( 0.0, 1.0] ( 0.0, 28.0+] ( 28.0, 308.0]
[25] ( 0.0, 36.0] ( 0.0, 20.0+] ( 20.0, 43.0]
[28] ( 0.0, 37.0] ( 0.0, 18.0+] ( 18.0, 28.0]
[31] ( 0.0, 8.0+] ( 8.0,1032.0] ( 0.0, 12.0+]
[34] ( 12.0, 51.0] ( 0.0, 3.0+] ( 3.0, 733.0]
[37] ( 0.0, 83.0+] ( 83.0, 219.0] ( 0.0, 25.0+]
[40] ( 25.0,1800.0+] ( 0.0,1401.0+] ( 0.0, 263.0]
[43] ( 0.0, 71.0+] ( 71.0, 72.0] ( 0.0, 35.0]
[46] ( 0.0, 16.0+] ( 16.0, 852.0] ( 0.0, 16.0]
[49] ( 0.0, 17.0+] ( 17.0, 77.0] ( 0.0, 51.0+]
[52] ( 51.0,1587.0+] ( 0.0, 23.0+] ( 23.0,1572.0+]
[55] ( 0.0, 12.0] ( 0.0, 46.0+] ( 46.0, 100.0]
[58] ( 0.0, 19.0+] ( 19.0, 66.0] ( 0.0, 4.5+]
[61] ( 4.5, 5.0] ( 0.0, 2.0+] ( 2.0, 53.0]
[64] ( 0.0, 41.0+] ( 41.0,1408.0+] ( 0.0, 58.0+]
[67] ( 58.0,1322.0+] ( 0.0, 3.0] ( 0.0, 2.0]
[70] ( 0.0, 40.0] ( 0.0, 1.0+] ( 1.0, 45.0]
[73] ( 0.0, 2.0+] ( 2.0, 996.0] ( 0.0, 21.0+]
[76] ( 21.0, 72.0] ( 0.0, 9.0] ( 0.0, 36.0+]
[79] ( 36.0,1142.0+] ( 0.0, 83.0+] ( 83.0, 980.0]
[82] ( 0.0, 32.0+] ( 32.0, 285.0] ( 0.0, 102.0]
[85] ( 0.0, 41.0+] ( 41.0, 188.0] ( 0.0, 3.0]
[88] ( 0.0, 10.0+] ( 10.0, 61.0] ( 0.0, 67.0+]
[91] ( 67.0, 942.0+] ( 0.0, 149.0] ( 0.0, 21.0+]
[94] ( 21.0, 343.0] ( 0.0, 78.0+] ( 78.0, 916.0+]
[97] ( 0.0, 3.0+] ( 3.0, 68.0] ( 0.0, 2.0]
[100] ( 0.0, 69.0] ( 0.0, 27.0+] ( 27.0, 842.0+]
[103] ( 0.0, 33.0+] ( 33.0, 584.0] ( 0.0, 12.0+]
[106] ( 12.0, 78.0] ( 0.0, 32.0] ( 0.0, 57.0+]
[109] ( 57.0, 285.0] ( 0.0, 3.0+] ( 3.0, 68.0]
[112] ( 0.0, 10.0+] ( 10.0, 670.0+] ( 0.0, 5.0+]
[115] ( 5.0, 30.0] ( 0.0, 31.0+] ( 31.0, 620.0+]
[118] ( 0.0, 4.0+] ( 4.0, 596.0+] ( 0.0, 27.0+]
[121] ( 27.0, 90.0] ( 0.0, 5.0+] ( 5.0, 17.0]
[124] ( 0.0, 2.0] ( 0.0, 46.0+] ( 46.0, 545.0+]
[127] ( 0.0, 21.0] ( 0.0, 210.0+] (210.0, 515.0+]
[130] ( 0.0, 67.0+] ( 67.0, 96.0] ( 0.0, 26.0+]
[133] ( 26.0, 482.0+] ( 0.0, 6.0+] ( 6.0, 445.0+]
[136] ( 0.0, 428.0+] ( 0.0, 32.0+] ( 32.0, 80.0]
[139] ( 0.0, 37.0+] ( 37.0, 334.0] ( 0.0, 5.0]
[142] ( 0.0, 8.0+] ( 8.0, 397.0+] ( 0.0, 60.0+]
[145] ( 60.0, 110.0] ( 0.0, 31.0+] ( 31.0, 370.0+]
[148] ( 0.0, 139.0+] (139.0, 207.0] ( 0.0, 160.0+]
[151] (160.0, 186.0] ( 0.0, 340.0] ( 0.0, 310.0+]
[154] (310.0, 340.0+] ( 0.0, 28.0+] ( 28.0, 265.0+]
[157] ( 0.0, 4.0+] ( 4.0, 165.0] ( 0.0, 2.0+]
[160] ( 2.0, 16.0] ( 0.0, 13.0+] ( 13.0, 180.0+]
[163] ( 0.0, 21.0+] ( 21.0, 131.0+] ( 0.0, 96.0+]
[166] ( 96.0, 109.0+] ( 0.0, 21.0] ( 0.0, 38.0+]
[169] ( 38.0, 39.0+] ( 0.0, 31.0+] ( 0.0, 11.0+]
[172] ( 0.0, 6.0]
2.survreg
拟合参数生存回归模型。 这些是用于时间变量的任意变换的位置尺度模型; 最常见的情况使用对数变换,建立加速失效时间模型。
survreg(formula, data, weights, subset,
na.action, dist="weibull", init=NULL, scale=0,
control,parms=NULL,model=FALSE, x=FALSE,
y=TRUE, robust=FALSE, score=FALSE, ...)
dist
y变量的假设分布。"weibull", "exponential", "gaussian", "logistic","lognormal" and "loglogistic"。
scale
可选的固定值。如果设置<=0,scale将被估计
# survreg's scale = 1/(rweibull shape) # survreg's intercept = log(rweibull scale) # survreg结果中输出的scale与“rweibull scale”不同 control
控制值列表,参考survreg.control()
survreg.control
survreg.control(maxiter=30, rel.tolerance=1e-09,
toler.chol=1e-10, iter.max, debug=0, outer.max=10)
maxiter
最大迭代次数
rel.tolerance
“相对容忍度”来声明收敛
toler.chol
Tolerance to declare Cholesky decomposition singular
iter.max
与maxiter相同
debug
打印调试信息
outer.max
用于选择惩罚参数的外部迭代的最大数目
convergence tolerance
/x(k+1)/=/x(k)/*Tr+Ta
其中:
k 迭代次数;
x(k+1) x的k次迭代计算值;
x(k) x的k次迭代初始值;
Tr 相对误差
Ta绝对误差
// 表示绝对值
因此,从上述公式我们可以得到一个重要结论:设定计算迭代误差的时候,要全面权衡物理量的绝对值大小,同时要衡量收敛迭代值的相对误差,这样才能正确满意的设定自己需要的计算误差!
不能简单的以为绝对误差和相对误差越小越好,也要杜绝tolerance设定时的随意性(按照公式进行合理的设定是一个优秀的过程工程师的基本素质)
survreg使用示例
> library(survival)
> str(ovarian)
'data.frame': 26 obs. of 6 variables:
$ futime : num 59 115 156 421 431 448 464 475 477 563 ...
$ fustat : num 1 1 1 0 1 0 1 1 0 1 ...
$ age : num 72.3 74.5 66.5 53.4 50.3 ...
$ resid.ds: num 2 2 2 2 2 1 2 2 2 1 ...
$ rx : num 1 1 1 2 1 1 2 2 1 2 ...
$ ecog.ps : num 1 1 2 1 1 2 2 2 1 2 ... > # 拟合指数模型,以下2个模型是一样的
> survreg(Surv(futime, fustat) ~ ecog.ps + rx, ovarian, dist='weibull',
+ scale=1)
Call:
survreg(formula = Surv(futime, fustat) ~ ecog.ps + rx, data = ovarian,
dist = "weibull", scale = 1) Coefficients:
(Intercept) ecog.ps rx
6.9618376 -0.4331347 0.5815027 Scale fixed at 1 # 注意这里 Loglik(model)= -97.2 Loglik(intercept only)= -98
Chisq= 1.67 on 2 degrees of freedom, p= 0.43
n= 26 > survreg(Surv(futime, fustat) ~ ecog.ps + rx, ovarian,
+ dist="exponential")
Call:
survreg(formula = Surv(futime, fustat) ~ ecog.ps + rx, data = ovarian,
dist = "exponential") Coefficients:
(Intercept) ecog.ps rx
6.9618376 -0.4331347 0.5815027 Scale fixed at 1 # 注意这里 Loglik(model)= -97.2 Loglik(intercept only)= -98
Chisq= 1.67 on 2 degrees of freedom, p= 0.43
n= 26 > ##########################################
>
> str(lung)
'data.frame': 228 obs. of 10 variables:
$ inst : num 3 3 3 5 1 12 7 11 1 7 ...
$ time : num 306 455 1010 210 883 ...
$ status : num 2 2 1 2 2 1 2 2 2 2 ...
$ age : num 74 68 56 57 60 74 68 71 53 61 ...
$ sex : num 1 1 1 1 1 1 2 2 1 1 ...
$ ph.ecog : num 1 0 0 1 0 1 2 2 1 2 ...
$ ph.karno : num 90 90 90 90 100 50 70 60 70 70 ...
$ pat.karno: num 100 90 90 60 90 80 60 80 80 70 ...
$ meal.cal : num 1175 1225 NA 1150 NA ...
$ wt.loss : num NA 15 15 11 0 0 10 1 16 34 ... > # weibull分布,如果设置scale<=0,模型将使用最大似然估计,估计最优的scale
> myfit<-survreg(Surv(time, status) ~ ph.ecog + age + sex, data=lung,dist = "weibull")
> myfit
Call:
survreg(formula = Surv(time, status) ~ ph.ecog + age + sex, data = lung,
dist = "weibull") Coefficients:
(Intercept) ph.ecog age sex
6.273435252 -0.339638098 -0.007475439 0.401090541 Scale= 0.731109 Loglik(model)= -1132.4 Loglik(intercept only)= -1147.4
Chisq= 29.98 on 3 degrees of freedom, p= 1.4e-06
n=227 (1 observation deleted due to missingness) > summary(myfit)
Call:
survreg(formula = Surv(time, status) ~ ph.ecog + age + sex, data = lung,
dist = "weibull")
Value Std. Error z p
(Intercept) 6.27344 0.45358 13.83 1.66e-43
ph.ecog -0.33964 0.08348 -4.07 4.73e-05
age -0.00748 0.00676 -1.11 2.69e-01
sex 0.40109 0.12373 3.24 1.19e-03
Log(scale) -0.31319 0.06135 -5.11 3.30e-07 Scale= 0.731 Weibull distribution
Loglik(model)= -1132.4 Loglik(intercept only)= -1147.4
Chisq= 29.98 on 3 degrees of freedom, p= 1.4e-06
Number of Newton-Raphson Iterations: 5
n=227 (1 observation deleted due to missingness) # survreg's scale = 1/(rweibull shape)
# survreg's intercept = log(rweibull scale)
# survreg结果中输出的scale与“rweibull scale”不同 ##############################################
> # weibull分布,以sex对数据分组,产生2个不同的scale
> myfit1<-survreg(Surv(time, status) ~ ph.ecog + age + strata(sex), data=lung,dist = "weibull")
> summary(myfit1)
Call:
survreg(formula = Surv(time, status) ~ ph.ecog + age + strata(sex),
data = lung, dist = "weibull")
Value Std. Error z p
(Intercept) 6.73235 0.42396 15.880 8.75e-57
ph.ecog -0.32443 0.08649 -3.751 1.76e-04
age -0.00581 0.00693 -0.838 4.02e-01
sex=1 -0.24408 0.07920 -3.082 2.06e-03
sex=2 -0.42345 0.10669 -3.969 7.22e-05 Scale:
sex=1 sex=2
0.783 0.655 Weibull distribution
Loglik(model)= -1137.3 Loglik(intercept only)= -1146.2
Chisq= 17.8 on 2 degrees of freedom, p= 0.00014
Number of Newton-Raphson Iterations: 5
n=227 (1 observation deleted due to missingness) ################################################
# 具有高斯误差的线性回归,以及左删失数据
> survreg(Surv(durable, durable>0, type='left') ~ age + quant,
+ data=tobin, dist='gaussian', scale = 0)
Call:
survreg(formula = Surv(durable, durable > 0, type = "left") ~
age + quant, data = tobin, dist = "gaussian", scale = 0) Coefficients:
(Intercept) age quant
15.14486636 -0.12905928 -0.04554166 Scale= 5.57254 Loglik(model)= -28.9 Loglik(intercept only)= -29.5
Chisq= 1.1 on 2 degrees of freedom, p= 0.58
n= 20
3.survdiff
测试在两条或更多条生存曲线之间是否存在差异
survdiff(formula, data, subset, na.action, rho=0)
rho = 0 表示log-rank or Mantel-Haenszel检验;
rho = 1 表示Wilcoxon检验
log-rank检验(时序检验)--生存时间分布近似weibull分布或者属于比例风险模型时效率较高
似然比检验(likelihood ratio test)--生存时间分布近似指数分布时效率较高
wilcoxon检验--生存时间分布近似对数正态分布效率较高
survdiff使用示例
> survdiff(Surv(futime, fustat) ~ rx,data=ovarian)
Call:
survdiff(formula = Surv(futime, fustat) ~ rx, data = ovarian) N Observed Expected (O-E)^2/E (O-E)^2/V
rx=1 13 7 5.23 0.596 1.06
rx=2 13 5 6.77 0.461 1.06 Chisq= 1.1 on 1 degrees of freedom, p= 0.303 > # 用strata来控制协变量的影响
> survdiff(Surv(time, status) ~ pat.karno + strata(inst), data=lung)
Call:
survdiff(formula = Surv(time, status) ~ pat.karno + strata(inst),
data = lung) n=224, 4 observations deleted due to missingness. N Observed Expected (O-E)^2/E (O-E)^2/V
pat.karno=30 2 1 0.692 0.13720 0.15752
pat.karno=40 2 1 1.099 0.00889 0.00973
pat.karno=50 4 4 1.166 6.88314 7.45359
pat.karno=60 30 27 16.298 7.02790 9.57333
pat.karno=70 41 31 26.358 0.81742 1.14774
pat.karno=80 50 38 41.938 0.36978 0.60032
pat.karno=90 60 38 47.242 1.80800 3.23078
pat.karno=100 35 21 26.207 1.03451 1.44067 Chisq= 21.4 on 7 degrees of freedom, p= 0.00326 > survdiff(Surv(time, status) ~ pat.karno + sex, data=lung)
Call:
survdiff(formula = Surv(time, status) ~ pat.karno + sex, data = lung) n=225, 3 observations deleted due to missingness. N Observed Expected (O-E)^2/E (O-E)^2/V
pat.karno=30, sex=1 1 1 0.246 2.31e+00 2.33e+00
pat.karno=30, sex=2 1 0 0.412 4.12e-01 4.16e-01
pat.karno=40, sex=1 1 1 0.132 5.68e+00 5.72e+00
pat.karno=40, sex=2 1 0 1.204 1.20e+00 1.22e+00
pat.karno=50, sex=1 2 2 0.111 3.23e+01 3.25e+01
pat.karno=50, sex=2 2 2 0.968 1.10e+00 1.11e+00
pat.karno=60, sex=1 18 17 9.173 6.68e+00 7.14e+00
pat.karno=60, sex=2 12 10 6.064 2.56e+00 2.68e+00
pat.karno=70, sex=1 30 23 16.737 2.34e+00 2.65e+00
pat.karno=70, sex=2 11 8 9.527 2.45e-01 2.62e-01
pat.karno=80, sex=1 32 26 24.570 8.32e-02 9.91e-02
pat.karno=80, sex=2 19 13 16.311 6.72e-01 7.55e-01
pat.karno=90, sex=1 31 25 24.992 2.66e-06 3.21e-06
pat.karno=90, sex=2 29 13 24.420 5.34e+00 6.33e+00
pat.karno=100, sex=1 21 15 14.002 7.11e-02 7.91e-02
pat.karno=100, sex=2 14 6 13.131 3.87e+00 4.25e+00 Chisq= 66.1 on 15 degrees of freedom, p= 2.17e-08
anova
计算一个或多个拟合模型的方差(或偏差)表的分析
mod1<-survreg(Surv(time, status) ~ ph.ecog + age + sex, data=lung,dist = "weibull")
mod2<-survreg(Surv(time, status) ~ ph.ecog + age + sex+ph.karno*pat.karno, data=lung,dist = "weibull") anova(mod1,mod2,test="Chisq")
Terms Resid. Df -2*LL Test Df Deviance Pr(>Chi)
1 ph.ecog + age + sex 222 2264.877 NA NA NA
2 ph.ecog + age + sex + ph.karno * pat.karno 215 2209.372 = 7 55.50527 1.183848e-09 library(MASS)
# 简洁模型更好
stepAIC(mod1)
stepAIC(mod2)
R生存分析AFT的更多相关文章
- R|生存分析 - KM曲线 ,值得拥有姓名和颜值
本文首发于“生信补给站”:https://mp.weixin.qq.com/s/lpkWwrLNtkLH8QA75X5STw 生存分析作为分析疾病/癌症预后的出镜频率超高的分析手段,而其结果展示的KM ...
- 生存分析与R
生存分析与R 2018年05月19日 19:55:06 走在码农路上的医学狗 阅读数:4399更多 个人分类: R语言 版权声明:本文为博主原创文章,未经博主允许不得转载. https://blo ...
- 生存分析与R--转载
生存分析与R 生存分析是将事件的结果和出现这一结果所经历的时间结合起来分析的一类统计分析方法.不仅考虑事件是否出现,而且还考虑事件出现的时间长短,因此这类方法也被称为事件时间分析(time-to-ev ...
- Forest plot(森林图) | Cox生存分析可视化
本文首发于“生信补给站”微信公众号,https://mp.weixin.qq.com/s/2W1W-8JKTM4S4nml3VF51w 更多关于R语言,ggplot2绘图,生信分析的内容,敬请关注小号 ...
- survival analysis 生存分析与R 语言示例 入门篇
原创博客,未经允许,不得转载. 生存分析,survival analysis,顾名思义是用来研究个体的存活概率与时间的关系.例如研究病人感染了病毒后,多长时间会死亡:工作的机器多长时间会发生崩溃等. ...
- R语言学习 - 非参数法生存分析--转载
生存分析指根据试验或调查得到的数据对生物或人的生存时间进行分析和推断,研究生存时间和结局与众多影响因素间关系及其程度大小的方法,也称生存率分析或存活率分析.常用于肿瘤等疾病的标志物筛选.疗效及预后的考 ...
- R数据分析:生存分析与有竞争事件的生存分析的做法和解释
今天被粉丝发的文章给难住了,又偷偷去学习了一下竞争风险模型,想起之前写的关于竞争风险模型的做法,真的都是皮毛哟,大家见笑了.想着就顺便把所有的生存分析的知识和R语言的做法和论文报告方法都给大家梳理一遍 ...
- Spark2 生存分析Survival regression
在spark.ml中,实现了加速失效时间(AFT)模型,这是一个用于检查数据的参数生存回归模型. 它描述了生存时间对数的模型,因此它通常被称为生存分析的对数线性模型. 不同于为相同目的设计的比例风险模 ...
- Cox回归模型【生存分析】
参考:<复杂数据统计方法--基于R的应用> 吴喜之 在生存分析中,研究的主要对象是寿命超过某一时间的概率.还可以描述其他一些事情发生的概率,例如产品的失效.出狱犯人第一次犯罪.失业人员第一 ...
随机推荐
- Python中使用MongoEngine
pymongo来操作MongoDB数据库,但是直接把对于数据库的操作代码都写在脚本中,这会让应用的代码耦合性太强,而且不利于代码的优化管理 一般应用都是使用MVC框架来设计的,为了更好地维持MVC结构 ...
- 常用Linq示例代码
class Program { static void Main(string[] args) { //1. Aggregate int[] testArr = new int[] { 1, 2, ...
- 详解BarTender符号体系特殊选项之“行数”
上面两篇文章小编和大家分享了BarTender符号体系特殊选项中的“行高”和“列”.此外,某些二维 (2D) 符号体系的结构为多个信息行,每一行看上去都像一个非常窄的条形码. 例如,以下图像是含 3 ...
- 求字符串长度StringLength();
// StringLength2.cpp : 定义控制台应用程序的入口点. // #include "stdafx.h" int StringLength(char str[]) ...
- json_encode让URL内容斜杠/不转义
同事在开发接口的时候根据接口提示要求传参一个字符串json,该json格式中有URL数组,按照json_encode编码后总发现 http://变成了 http:\/\/ .URL的斜杠自动的被转义 ...
- 转载nginx+uwsgi+django
Django的部署可以有很多方式,采用nginx+uwsgi的方式是其中比较常见的一种方式. 在这种方式中,我们的通常做法是,将nginx作为服务器最前端,它将接收WEB的所有请求,统一管理请求.ng ...
- 数组名和数组名取地址&
在C中, 在几乎所有使用数组的表达式中,数组名的值是个指针常量,也就是数组第一个元素的地址. 它的类型取决于数组元素的类型: 如果它们是int类型,那么数组名的类型就是“指向int的常量指针“ ...
- /var/spool/postfix/maildrop/ 中有大量的文件
今天查看硬盘剩余的容量,发现‘/’目录下占用了大量的空间:可我在这个目录下面没有放什么东西:仔细查看在/var/spool/postfix/maildrop/ 中发现了大量的文件.怎么会有这么多的文件 ...
- Linux下chkconfig命令详解转载
chkconfig命令主要用来更新(启动或停止)和查询系统服务的运行级信息.谨记chkconfig不是立即自动禁止或激活一个服务,它只是简单的改变了符号连接. 使用语法:chkconfig [--ad ...
- Lua中的控制结构
Lua提供了一组传统的.小巧的控制结构,包括用于条件执行的if,用于迭代的while.repeat和for.所有的控制结构都有意个显式的终止符:if.for和while以end作为结尾,repeat以 ...
